Non-Invasive Prenatal Testing (NIPT), also known as cell-free DNA prenatal screening, is a widely adopted clinical technology for detecting fetal chromosomal aneuploidies. By analyzing cell-free fetal DNA in maternal peripheral blood, this approach eliminates the risks associated with traditional invasive procedures such as amniocentesis and significantly improves the accuracy of detecting common chromosomal abnormalities. Fetal Fraction (FF), the proportion of cell-free fetal DNA in the total cell-free DNA within maternal plasma, is a critical quality metric for assessing the reliability of NIPT results.
In recent years, digital PCR has emerged as a promising platform for cell-free DNA prenatal screening due to its operational simplicity and relatively low cost. However, this platform currently lacks an FF quantification method that can be seamlessly integrated into its workflow. Existing FF determination methods generally suffer from limitations such as sex dependency, complex procedures, or the need for parental genotyping, which hinders their further clinical application and adoption.
To address these challenges, a collaborative research team led by Professor Guo Yong from the School of Biomedical Engineering at Tsinghua University, in partnership with the First Affiliated Hospital of Zhengzhou University, Hunan University of Medicine, Dongguan Maternal and Child Health Hospital, and TargetingOne Corporation, has developed a novel digital PCR method for FF quantification based on multiplex single nucleotide polymorphism (SNP) and Y chromosome (chrY) detection. Through an innovative single-probe, dual-allele discrimination design and a triplex Y chromosome target strategy, this method achieves accurate FF quantification without the need for parental genotyping and is independent of fetal sex. It provides a critical quality control tool for advancing the clinical translation of digital PCR-based cell-free DNA prenatal screening. The findings, titled "A Digital PCR Assay for Fetal Fraction Quantification Using Multiplex SNP and Y Chromosome Detection," were published in Clinical Chemistry on February 26, 2026.

Click for more information on the paper
Four Technological Innovations Addressing Current Limitations
1: Sex-Independent Dual-Track Detection Strategy: Simultaneously detects 9 autosomal SNP loci and 3 chrY loci. For male fetuses, the chrY method is used (offering higher precision); for female fetuses, the SNP method is applied.
2: Single-Probe, Dual-Allele Discrimination Technology: Utilizes an optimized single TaqMan probe that distinguishes two alleles simultaneously in a single fluorescence channel. Through precise thermodynamic control of probe melting temperature, SNP genotyping is achieved based solely on differences in fluorescence intensity.
3: Triplex chrY Targets Enhance Sensitivity: Targeting 3 distinct regions on the chrY increases the number of positive droplets at low concentrations, overcoming Poisson statistical noise and significantly improving detection accuracy in low-FF samples.
4: Algorithm Eliminates Need for Parental Genotyping: Automatically calculates FF using a biologically constrained algorithm (FF > 0% and < 50%), simplifying the clinical workflow.
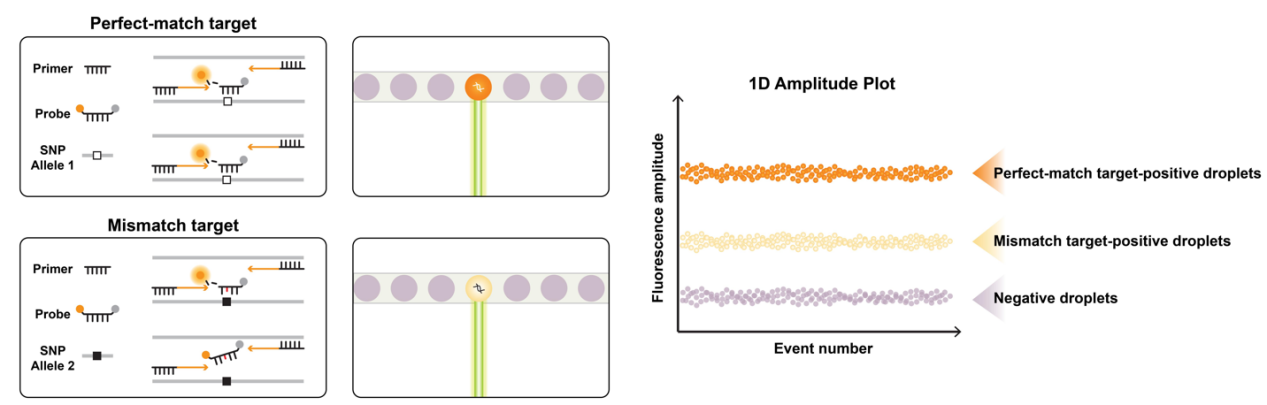
Single-Probe, Dual-Allele Discrimination Technology
Simple Three-Step Detection Workflow Compatible with Clinical Practice
1: Sample Processing and DNA Extraction: Collect maternal peripheral blood and extract cell-free DNA from the plasma.
2: Multiplex Digital PCR Detection: Use an automated digital PCR platform to detect 9 autosomal SNP loci and 3 chrY loci in two well reaction.
3: Data Analysis and FF Calculation: Differentiate allele types based on droplet fluorescence amplitude, calculate copy numbers for each target, apply limit of detection (LOD) threshold filtering and biological plausibility constraints, and output the final FF value.
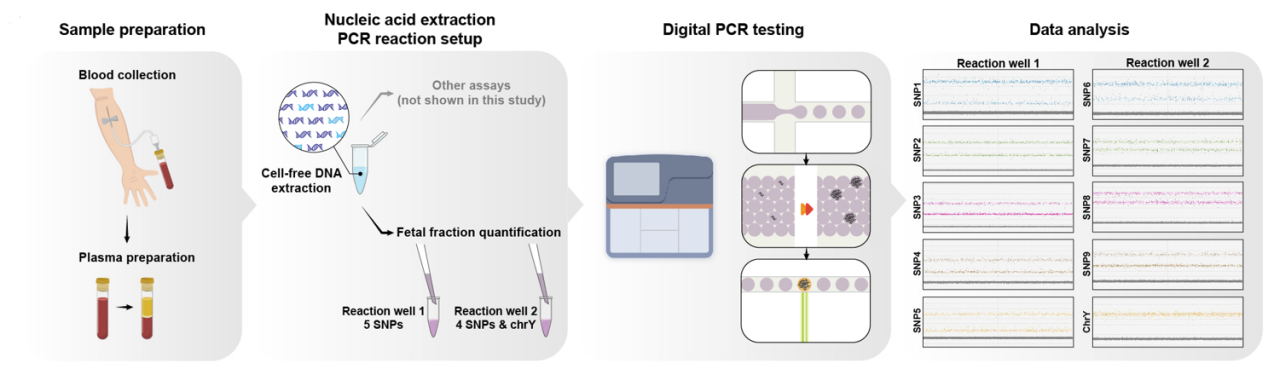
Overall Workflow
Performance Validation: Sensitive and Accurate
1: Excellent Detection Sensitivity: The chrY method achieves a minimum detectable FF of 1%, while the SNP method has a LOD of 2.5%.
2: Good Methodological Consistency: Validated with clinical samples, the results obtained with this method demonstrate high consistency with those from next-generation sequencing (NGS) methods.
3: Rapid Assay Turnaround Time: The complete detection process for a single sample takes approximately 3.5 hours.
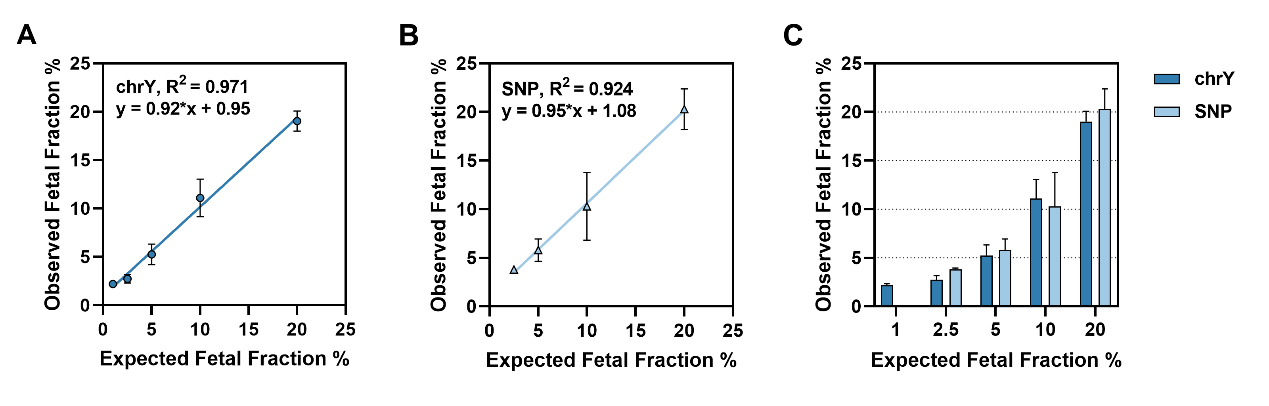
Analytical Validation Using Simulated Samples
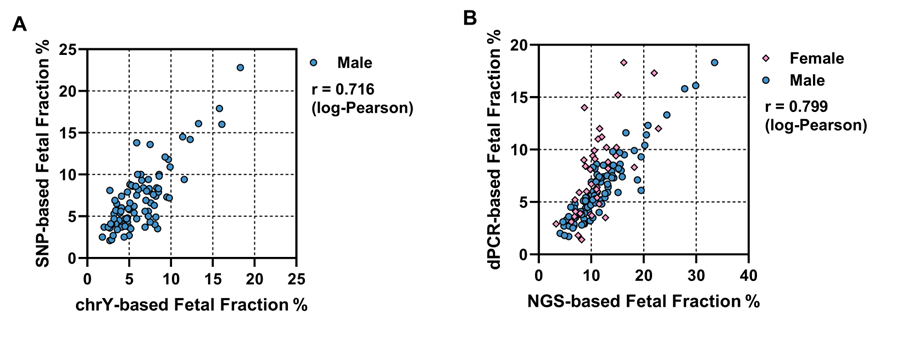
Clinical Validation: Consistency Assessment between Digital PCR and NGS Methods
Multidimensional Applications Supporting Birth Defect Prevention and Control
This technology offers significant clinical and social value through its multidimensional applications. In non-invasive prenatal screening, it can serve as a pre-analytical quality assessment tool for digital PCR tests, quickly determining whether sample FF meets testing requirements, effectively reducing the risk of false negatives, and improving detection accuracy. Compared to NGS, it offers a workflow with significantly lower technical complexity, faster turnaround, and reduced cost, making it more suitable as a rapid QC step in routine laboratories. Furthermore, this method shows potential for monitoring obstetric complications by dynamically tracking changes in maternal FF, aiding in the prediction of pregnancy-related conditions such as preeclampsia.
This work is part of a series of advancements by the team in the field of digital PCR-based NIPT. In 2019, the team published a cover article in Analyst demonstrating the use of multiplex digital PCR for non-invasive detection of fetal chromosomal aneuploidies. Building on that foundation, the team has continued to innovate, developing a comprehensive technical system that now includes both chromosomal aneuploidy detection and FF quantification. This integrated approach combines technical robustness with economic accessibility, playing a vital role in expanding the application of cell-free DNA prenatal screening and improving the coverage of birth defect prevention and control efforts.
Professor Guo Yong from the School of Biomedical Engineering, Tsinghua University, is the corresponding author of the paper. Dr. Tan Chianru (Malaysian nationality), a 2023 Ph.D. graduate from the same school and currently a "Shuimu Scholar" postdoctoral fellow, and Dr. Dai Peng from the First Affiliated Hospital of Zhengzhou University, are the co-first authors. The research was supported by grants including the Beijing Nova Program, the SXMU-Tsinghua Collaborative Innovation Center for Frontier Medicine, the Key Project of the Medical Science and Technology Research Program of Henan Province, and the Henan Provincial Science and Technology Research Project.


